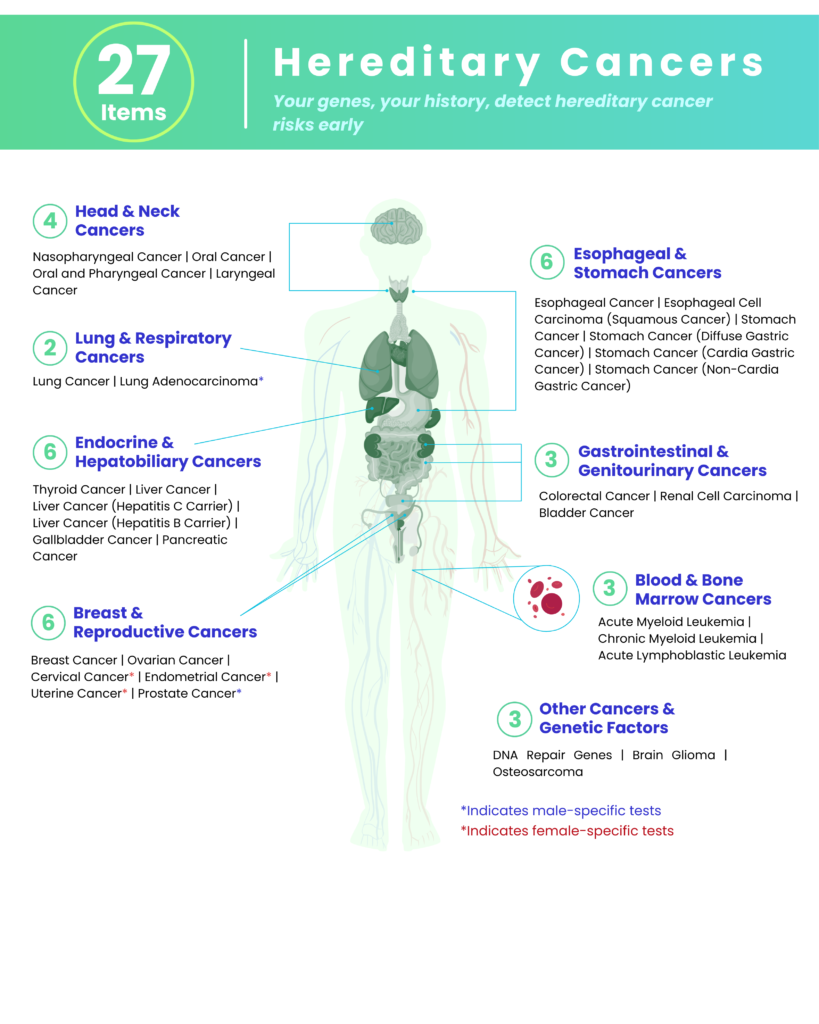
Your genes hold the key to your well-being, and Blueprint of Life™ empowers you with insights for a healthier future. Our comprehensive genetic test covers 94 health risk indicators (101 for women), helping you understand your unique genetic makeup, assess potential health risks, and make informed decisions about your health management.
With an extensive analysis spanning

Cancer

Disease Risks

Wellness & Nutrition

Beauty & Ageing

Hereditary Risks
Blueprint of Life™ provides a holistic view of your health. Our non-invasive genetic testing ensures a comfortable experience without compromising accuracy.
Receive a detailed, easy-to-understand report, coupled with expert consultation to guide you in personalizing your health strategies, whether for:
Prevention
Lifestyle management
Treatment decisions
How Does It Work?

❶ Sample Collection
Collecting DNA samples can be made comfortable and easy through non-invasive methods, such as using buccal cells obtained from mouthwash. This method offers:
- Comfort
Buccal cell collection is painless, making it ideal for individuals who are hesitant about invasive procedures - Convenience
The process involves simply rinsing the mouth with mouthwash solution and collecting it inside a sample tube provided - Cost-effectivity
This method is inexpensive compared to other invasive procedures, which require more specialized equipment and trained personnel - High-Quality DNA
Buccal cells can provide sufficiently high-quality DNA for genetic analysis
❷ Library Preparation
Sequencing Techniques Development
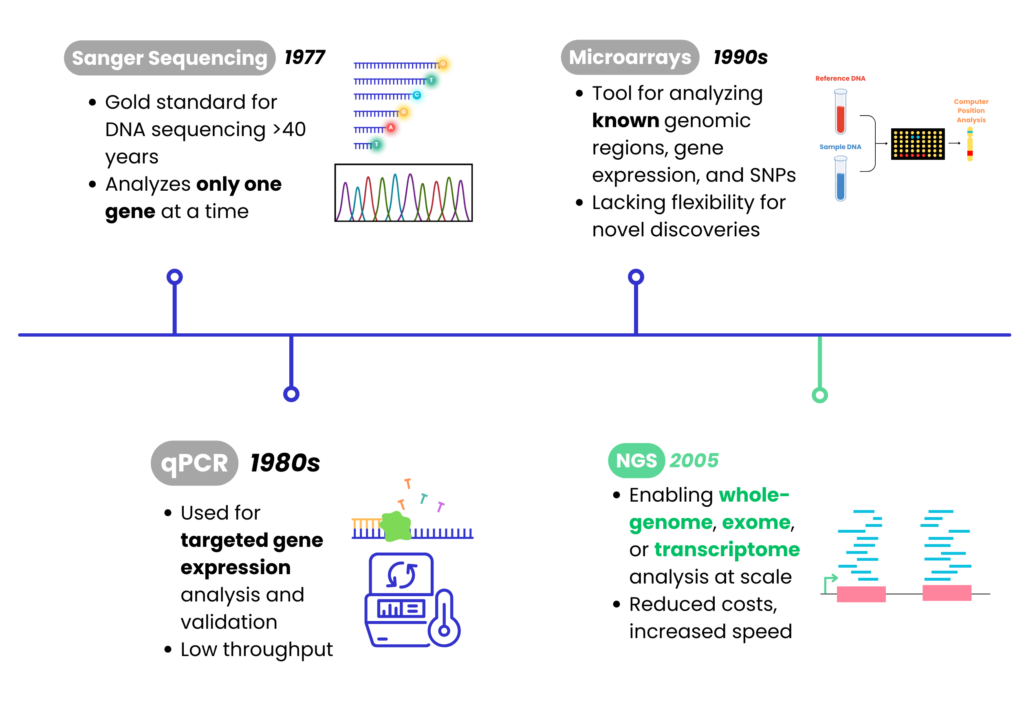
NGS vs. Microarray vs. qPCR Comparisons
NGS outperforms microarrays and qPCR in throughput, coverage, and discovery power, making it the preferred choice for modern genomics.
| Parameter | NGS (Next Generation Sequencing) | Microarray | qPCR |
| Detection Principle | Direct sequencing, analyzing DNA sequences | Probe hybridization, detecting specific gene variations | Uses fluorescent probes to monitor PCR-amplified DNA products in real-time |
| Detectable Variants | Point mutations (SNV), insertions/deletions (INDELs), copy number variations (CNV), structural variations | Known single nucleotide polymorphisms (SNP) or copy number variations (CNV) | Known single nucleotide polymorphisms (SNP), insertions/deletions (INDELs), gene expression level changes |
| Analysis | Bioinformatics analysis | Low, only detects pre-designed variations | High, but with limited mutation coverage |
| Applications | Non-invasive prenatal testing (NIPT), cancer gene testing, rare disease diagnosis | Genotyping chips, population research, Biobank | COVID-19 diagnosis, alcohol metabolism genes |
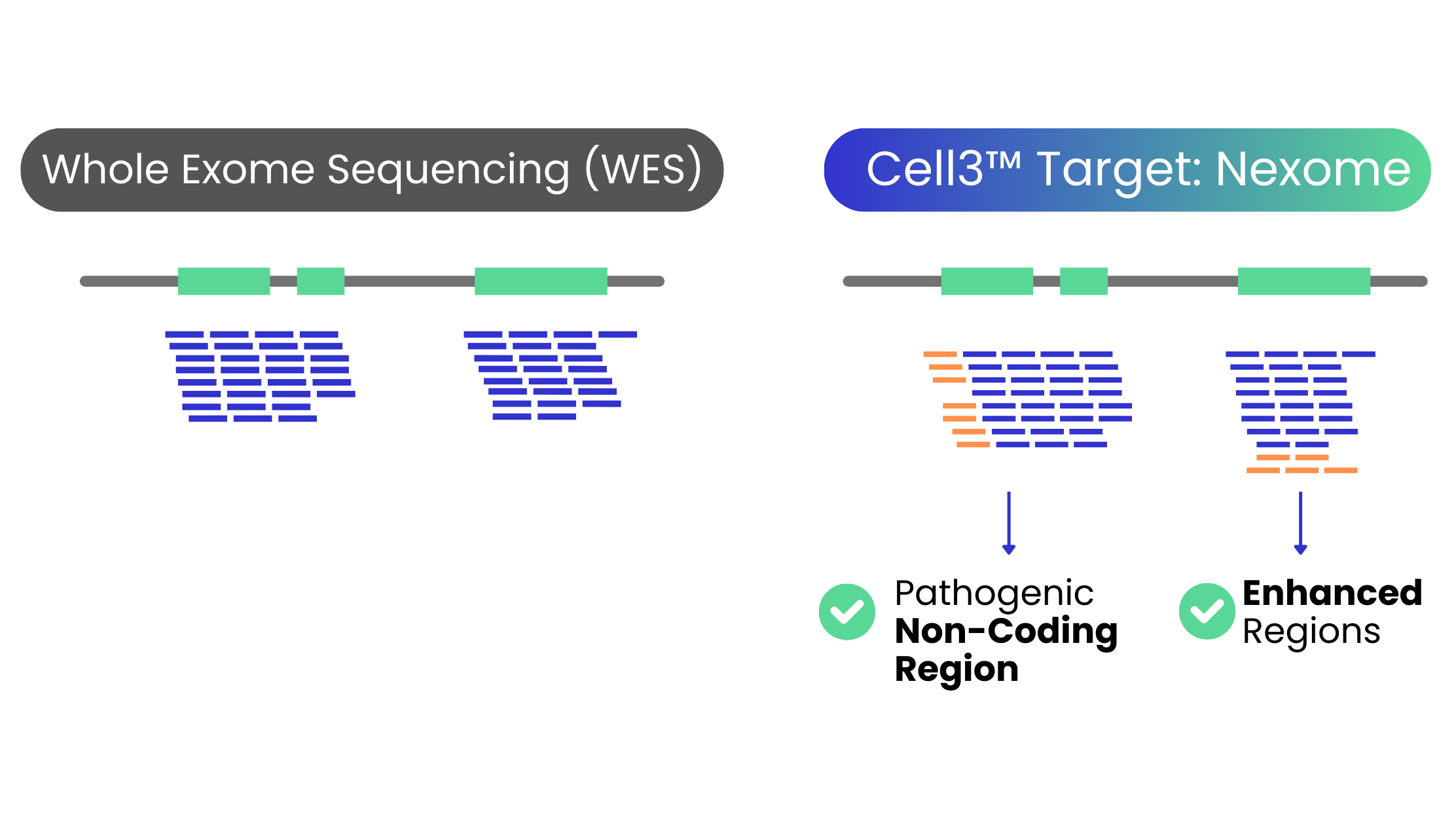
Cell3™ Target: Nexome 一 Advancing Beyond Traditional WES
Cell3™ Target: Nexome is a cutting-edge next-generation exon sequencing panel, designed to overcome the limitations of traditional whole-exome sequencing (WES).
By combining advanced probe design with superior coverage and uniformity, Cell3™ Target: Nexome offers a comprehensive and cost-effective solution for genomics and research applications.
The panel includes
Genes linked to prenatal diagnosis, epilepsy, and pharmacogenomics (PGx)

Exon-level deletions and duplications

Non-coding variants linked to genetic diseases

RefSeq transcripts, promoter regions, 5′ and 3′ UTR sequences corresponding to OMIM morbid set of 4,090 genes
Approximately 85% diseases caused by DNA mutations occur due to variations in the exon regions.
Cell3™ Target: Nexome panel captures up to 30% more variants than commercially available exome products
| Feature | Traditional WES | Cell3™ Target: Nexome |
| Genomic Coverage | Primarily focuses on protein-coding regions (1-2% of genome) | Protein-coding parts of the genome + clinically relevant non-coding regions |
| Type of Variants Detected | SNVs and INDELs but requires additional tests for CNVs | Single nucleotide variants (SNVs), insertions/deletions (INDELs), and copy number variations (CNVs) in one test |
| Diagnostic Yield | Limited to coding variants, may miss some clinically relevant variants | Detects up to 30% more variants than standard exome panels |
| Efficiency and Cost | Requires separate tests for CNVs, which can increase costs and complexity | Streamlines the diagnostic process by combining multiple tests into one, potentially reducing overall costs |
| Applications | General genetic diagnostics but may not offer the same level of comprehensive analysis | Prenatal diagnostics, epilepsy evaluation, and cases requiring comprehensive genetic analysis |
❸ Reporting
🎯 Covering Over 200+ Items
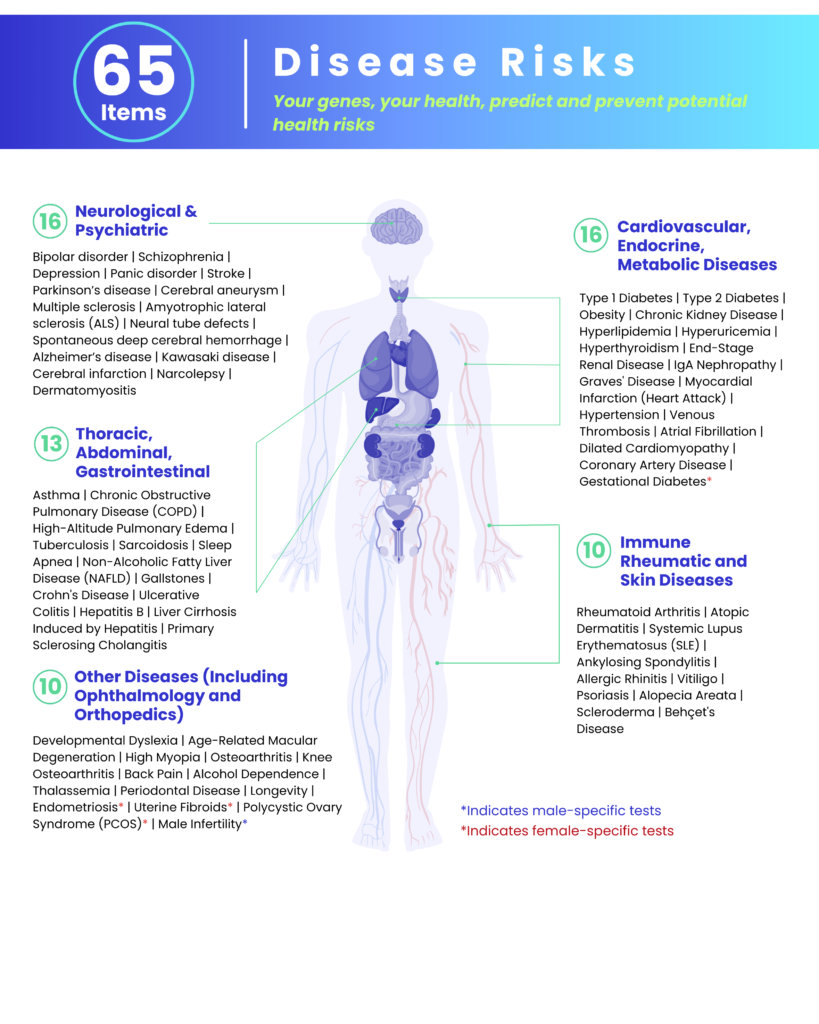
- 16 Neurological & Psychiatric
- 13 Thoracic, Abdominal, Gastrointestinal
- 16 Cardiovascular, Endocrine, Metabolic Diseases
- 10 Immune Rheumatic & Skin Diseases
- 10 Other Diseases (Including Ophthalmology & Orthopedics)
- 4 Head & Neck Cancers
- 2 Lung & Respiratory Cancers
- 6 Endocrine & Hepatobiliary Cancers
- 6 Breast & Reproductive Cancers
- 6 Esophageal & Stomach Cancers
- 3 Gastrointestinal & Genitourinary Cancers
- 3 Blood & Bone Marrow Cancers
- 3 Other Cancers & Genetic Factors
- 9 Physical Fitness & Constitution
- 7 Taste & Smell Sensitivity
- 4 Dietary Habits & Nutrient Absorption
- 2 Metabolism & Weight Control
- 14 Weight Loss Genes
- 13 Vitamin & Mineral Levels
(Blood Concentration) - 16 Personal Traits
- 11 Medical Aesthetics
- 14 Talents
- 10 Other Health Factors
- 6 Blood Disorders
- 26 Endocrine & Metabolic Disorders
- 11 Neurological & Muscular Disorders
- 4 Bone & Connective Tissue Disorders
- 6 Thyroid Disorders
- Other Genetic Disorders
Understand More about Disease Risks Genetic Test Items
Reports Benchmarked Against a Diverse Asian Population Database
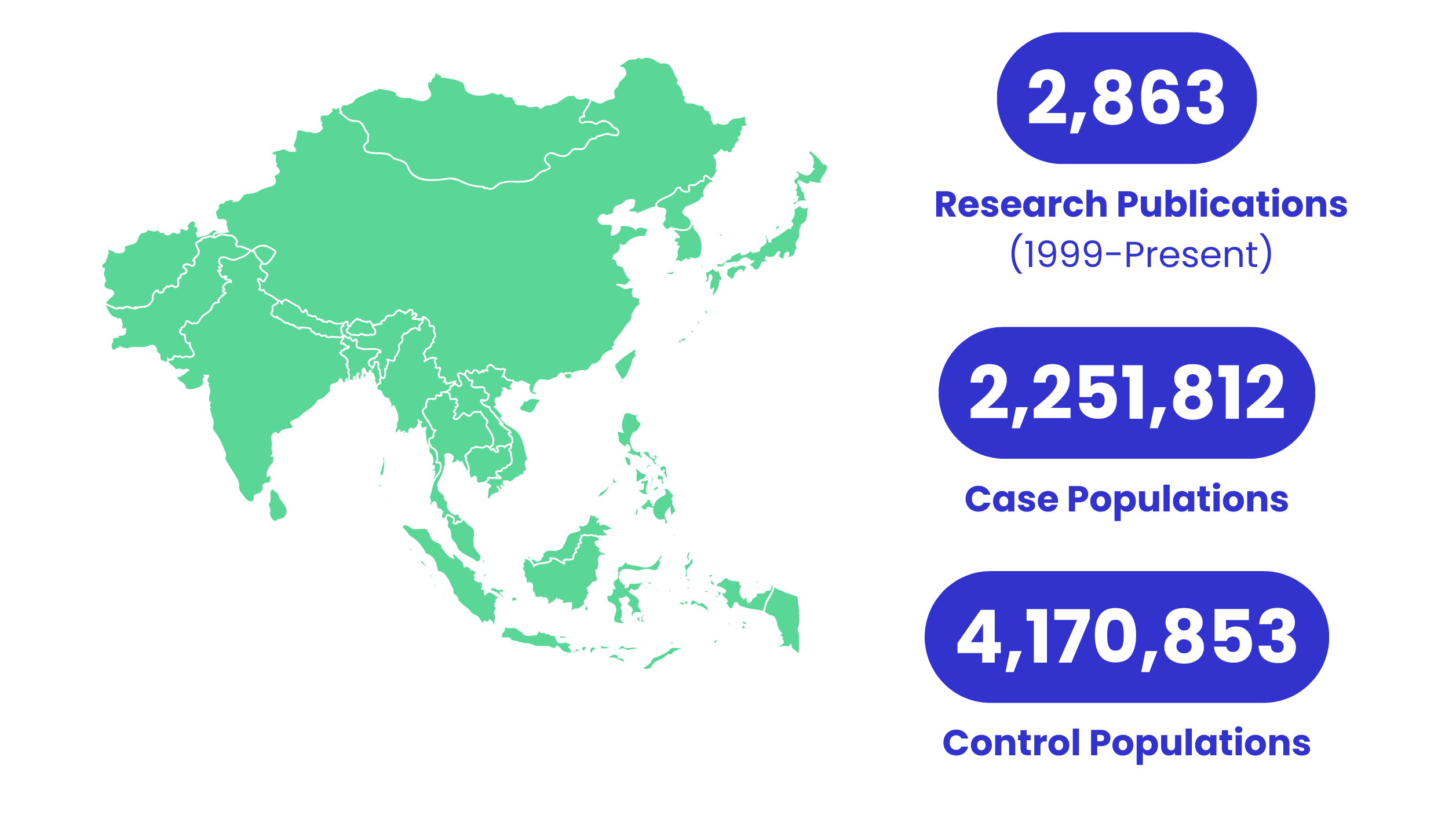
Our reports are meticulously benchmarked against a comprehensive Asian population database, ensuring that our genetic insights are tailored to the unique genetic profiles of Asian populations. This approach is crucial for providing accurate and relevant genetic information, particularly for individuals of Chinese descent.
We have curated a vast collection of research literature from global databases, focusing on studies relevant to Asian genetic types. This extensive compilation spans 2,863 research articles published between 1999 and the present, encompassing data from 2,251,812 case populations and 4,170,853 control populations.
By using this diverse Asian population database, we enhance the accuracy and relevance of our genetic reports. This approach helps identify genetic factors that are particularly significant for Asian populations, contributing to more personalized and effective healthcare strategies.
Accredited Laboratory Excellence: Internationally Certified
LDTS
ISO 15189
TAF

❹ After Service
Post-Test Support: Genetic Counseling & Expert Guidance
After receiving your genetic test results, understanding the implications and making informed decisions can be challenging. Our post-test support ensures that you have access to expert guidance through consultations with genetic counselors and specialists. This comprehensive support helps you navigate the complexities of genetic testing, ensuring that you fully understand your results and their implications for your health and family.
Informed Decision-Making
Helps you make informed decisions about your health and family planning based on your genetic information
Personalized Care
Facilitates personalized health management by discussing potential treatment options and referrals to specialists
References
- Nature news. https://www.nature.com/scitable/topicpage/inheritance-of-traits-by-offspring-follows-predictable-6524925/
- Loewe, L. (2008) Genetic mutation. Nature Education 1(1):113
- Stallard, T. (2023) Rare disease care across asia pacific, Sandpiper.
- Hlawulani (2019) New Scientific Paper confirms 300 million people living with a rare disease worldwide, Rare Diseases International.
- Pesheva, E. (2023) Researchers able to determine the effects of genes and environment in 560 common conditions, Harvard Gazette.
- Hanany, M., Rivolta, C., & Sharon, D. (2020). Worldwide carrier frequency and genetic prevalence of autosomal recessive inherited retinal diseases. Proceedings of the National Academy of Sciences, 117(5), 2710–2716.
- Abdul, Q. A., Yu, B. P., Chung, H. Y., Jung, H. A., & Choi, J. S. (2017). Epigenetic modifications of gene expression by lifestyle and environment. Archives of Pharmacal Research, 40(11), 1219–1237. https://doi.org/10.1007/s12272-017-0973-3
- Alegría-Torres, J. A., Baccarelli, A., & Bollati, V. (2011). Epigenetics and Lifestyle. Epigenomics, 3(3), 267–277.
- CDC. (2025, January 31). Epigenetics, Health, and Disease. Genomics and Your Health. https://www.cdc.gov/genomics-and-health/epigenetics/index.html
- Advantages of NGS Over Other Molecular Methods. (2020). Illumina.com. https://sapac.illumina.com/science/technology/next-generation-sequencing/beginners/advantages.html
- UF scientists test mouthwash method of collecting DNA – UF Health. (2023). Ufhealth.org. https://ufhealth.org/news/2002/uf-scientists-test-mouthwash-method-collecting-dna
- Zayats, T., Young, T. L., Mackey, D. A., Malecaze, F., Calvas, P., & Guggenheim, J. A. (2009). Quality of DNA Extracted from Mouthwashes. PLoS ONE, 4(7). https://doi.org/10.1371/journal.pone.0006165
- Lelieveld, S. H., Spielmann, M., Mundlos, S., Veltman, J. A., & Gilissen, C. (2015). Comparison of Exome and Genome Sequencing Technologies for the Complete Capture of Protein-Coding Regions. Human Mutation, 36(8), 815–822.
- Zhao, Y., Fang, L. T., Shen, T., Choudhari, S., Talsania, K., Chen, X., Shetty, J., Kriga, Y., Tran, B., Zhu, B., Chen, Z., Chen, W., Wang, C., Jaeger, E., Meerzaman, D., Lu, C., Idler, K., Ren, L., Zheng, Y., & Shi, L. (2021). Whole genome and exome sequencing reference datasets from a multi-center and cross-platform benchmark study. Scientific Data, 8(1). https://doi.org/10.1038/s41597-021-01077-5
- Barbitoff, Y. A., Polev, D. E., Glotov, A. S., Serebryakova, E. A., Shcherbakova, I. V., Kiselev, A. M., Kostareva, A. A., Glotov, O. S., & Predeus, A. V. (2020). Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage. Scientific Reports, 10(1). https://doi.org/10.1038/s41598-020-59026-y